 A program to define the genetic structure of populations by a simulated annealing approach
A program to define the genetic structure of populations by a simulated annealing approach  A program to define the genetic structure of populations by a simulated annealing approach
A program to define the genetic structure of populations by a simulated annealing approach SAMOVA 2.0 implements an approach to define groups of populations that are geographically homogeneous and maximally differentiated from each other. As a by-product, it also leads to the identification of genetic barriers between these groups. The method is based on a simulated annealing procedure that aims at maximizing the proportion of total genetic variance due to differences between groups of populations (SAMOVA, Spatial Analysis of MOlecular VAriance). The method is described in Dupanloup, Schneider and Excoffier (2002).
A new functionality of SAMOVA 2.0 is to define groups of populations that are maximally differentiated from each other, without constraint for the geographic composition of the groups.
SAMOVA 2.0 runs on Windows. There is no Linux or Mac version yet.
Preliminary steps
Simulated annealing steps
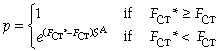
In SAMOVA 1.0, the number of steps S in the simulated annealing process was set to 10 000 and the constant A to 0.9158. In this case, the probability p defined above is equal to 1% if the difference between FCT and FCT* at the 10 000th iteration is equal to 0.001. In SAMOVA 2.0, S and A can be chosen by the user. You'll find below different combinations of values of S and A and the corresponding probability values to accept the new structure
| FCT*-FCT | S | A | p |
| -0.1 | 10000 | 0.915811421 | 1E-200 |
| -0.01 | 10000 | 0.915811421 | 1E-20 |
| -0.001 | 10000 | 0.915811421 | 0.01 |
| -0.1 | 100 | 1.831622842 | 1E-200 |
| -0.01 | 100 | 1.831622842 | 1E-20 |
| -0.001 | 100 | 1.831622842 | 0.01 |
| -0.1 | 1000 | 1.221081895 | 1E-200 |
| -0.01 | 1000 | 1.221081895 | 1E-20 |
| -0.001 | 1000 | 1.221081895 | 0.01 |
| -0.1 | 100000 | 0.732649137 | 1E-200 |
| -0.01 | 100000 | 0.732649137 | 1E-20 |
| -0.001 | 100000 | 0.732649137 | 0.01 |
| -0.1 | 1000000 | 0.610540947 | 1E-200 |
| -0.01 | 1000000 | 0.610540947 | 1E-20 |
| -0.001 | 1000000 | 0.610540947 | 0.01 |
To make sure that the final configuration of the K groups is not affected by a given initial configuration, the simulated annealing process is repeated 100 times, starting each time from a different initial partition of the n samples into the K groups. The configuration with the largest associated FCT value after the 100 independent simulated annealing processes is retained as the best grouping of populations.
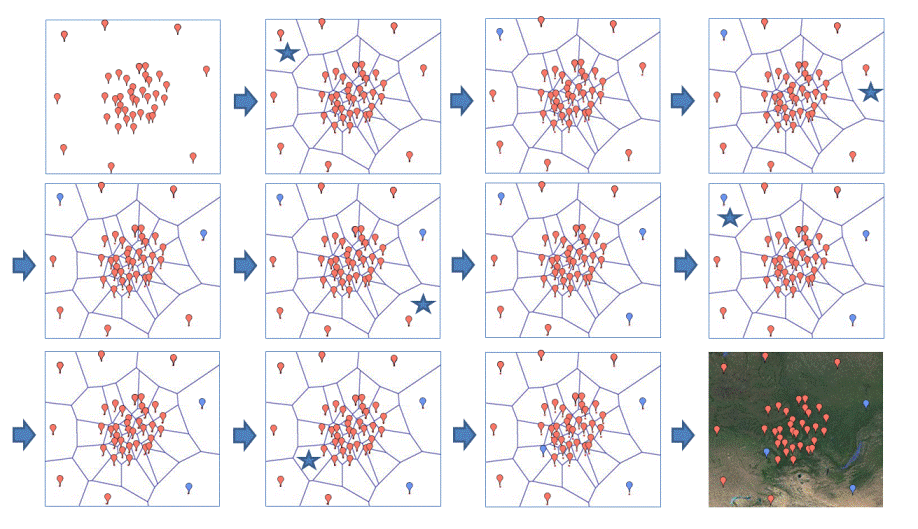
The cartoon below illustrates the behaviour of SAMOVA 2.0.
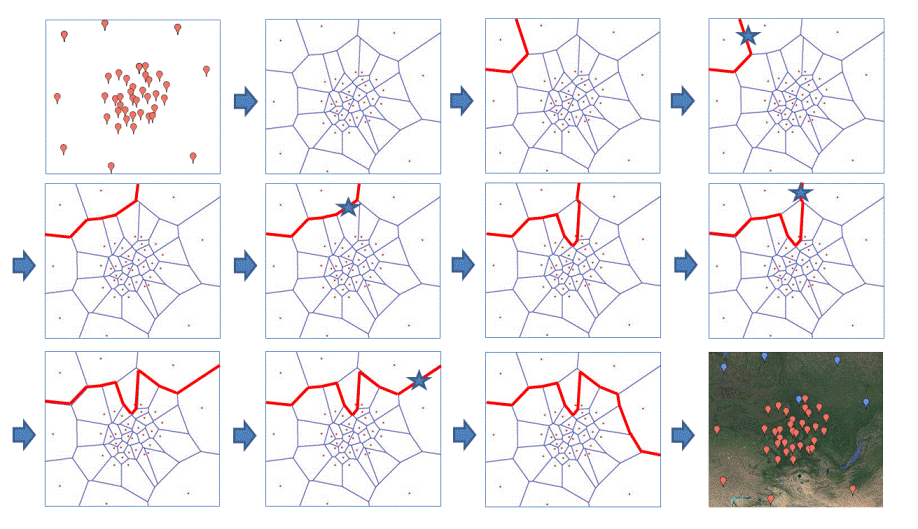
The cartoon below illustrates one case encountered frequently with SAMOVA 2.0: the allocation of one population from one group to another leads to the fragmentation of one group in 2 distinct sets of adjacent populations.
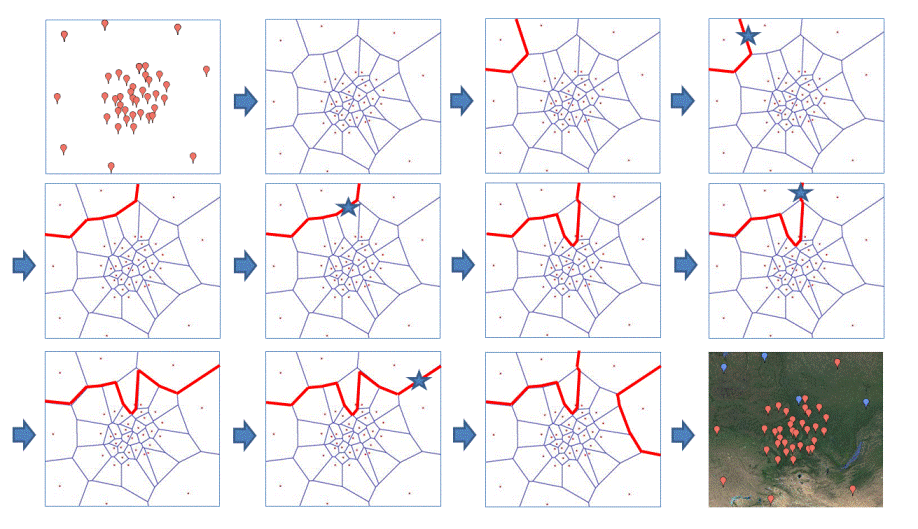
Preliminary steps
Simulated annealing steps
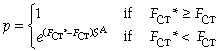
The cartoon below illustrates the behaviour of SAMOVA 2.0.
SAMOVA 2.0 (like SAMOVA 1.0) needs two input files. The first one (*.geo) must contain the geographic coordinates of the sampling localities of your populations. The second one (*.arp) is an Arlequin input file containing the genetic data sampled in your populations. The Arlequin file must have the SAME NAME as the geographical file with the extension (*.arp). The order of the populations in the two input files MUST BE THE SAME !!!
The file containing the geographic coordinates of the sampling localities of your populations must have the .geo extension.
Important notice: SAMOVA 2.0 does not work if two sampling localities have the same geographical coordinates.
The geographical input file must be structured the following way. Each line corresponds to a population. Each line must contain five fields separated by TAB characters:
Examples of input files are given below:
When SAMOVA 2.0 runs, it expects the generic name of your input files. If you have INPUTFILE.GEO and INPUTFILE.ARP as input files, it will expect to read INPUTFILE (either in the INPUTFILE.SAR file or from the standard input).
A set of output files are created by SAMOVA:
Isabelle Dupanloup, CMPG, Institute of Ecology and Evolution, University of Bern